Global patterns emerge from microbial interactions
Recent sequencing surveys have revealed that global relationships between microbiome structure (taxonomy and gene content) and environmental factors (temperature, pH, nutrient availability, etc.) are ubiquitous. To explain these relationships, the concept of environmental filtering – that the environment “filters” the microbial community based upon which microbes are best adapted to a given environment – is often invoked. While environmental filtering provides a natural explanation for the global conservation of these environment-structure relationships (if the microbes present in an environment depend solely on that environment, the relationship should be highly conserved), it is difficult to reconcile with the extensive and well-documented impact of microbial interactions on microbiome content.
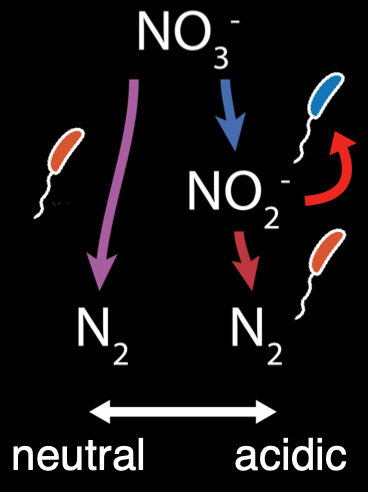
To resolve this paradox, I use a combination of statistics, experiments, and theory to understand the origin of global patterns in microbiome content. I first analyze a topsoil microbiome dataset (Bahram et al., 2018) and identify a relationship between genotype and pH. Next, I bring natural soil communities into the lab and show that this relationship can be recapitulated experimentally. Finally, I isolate the microbes driving this relationship and perform detailed experiments on these isolates. I show that, rather than arising from simple adaptation of microbe to environment, the pH-genotype relationship is dependent on interactions between microbes. In particular, community interactions that alleviate toxicity emerge in acidic environments. These interactions, therefore, are themselves modulated by the environment. This results in interactions that are conserved in similar environments, giving rise to the observed global patterns.
This work was carried out in collaboration with fellow postdoc Karna Gowda and graduate student Milena Chakraverti-Wuerthwein, and was advised by Profs. Seppe Kuehn and Madhav Mani.